Science Magazine's "Insights of the Decade"
Berkeley Lab's NERSC and Computational Research divisions contributed to four of the ten discoveries
January 31, 2011
By John Hules
Contact: cscomms@lbl.gov
In the first decade of this millennium, rapid scientific progress—resulting from new sensing, imaging, computing, and networking tools—has transformed whole areas of research. The December 17, 2010 special issue of Science magazine highlights ten “Insights of the Decade” that have fundamentally changed our thinking about our bodies and our universe. Four of those ten insights were enabled in part by facilities and research in Lawrence Berkeley National Laboratory’s (Berkeley Lab’s) National Energy Research Scientific Computing Center (NERSC) and Computational Research Division (CRD).
In their introduction to the special section, Science magazine’s news staff gives credit to the new tools that made these insights possible. They write:
"In the past 10 years, new ways of gathering, analyzing, storing, and disseminating information have transformed science. Researchers generate more observations, more models, and more automated experimentation than ever before, creating a data-saturated world. The Internet has changed how science is communicated and given nonscientists new opportunities to take part in research. Whole new fields, such as network science, are arising, and science itself is becoming more of a network—more collaborative, more multidisciplinary—as researchers recognize that it takes many minds and varied expertise to tackle complex questions about life, land, and the universe…."
Key to handling such unprecedented torrents of data, of course, have been ever more powerful and more affordable computers. No field has benefited more than genomics. A decade ago, sequencing a human genome took years, hundreds of people, hundreds of machines, and endless hours of sample preparation to generate the pieces of DNA to be deciphered, one at a time…. Today, a single machine can decipher three human genomes in little more than a week.
Three of the ten Insights of the Decade were in the field of genomics, and the U.S. Department of Energy (DOE) Joint Genome Institute (JGI), supported by NERSC and CRD, was involved in all three.
DNA's "Dark Matter"
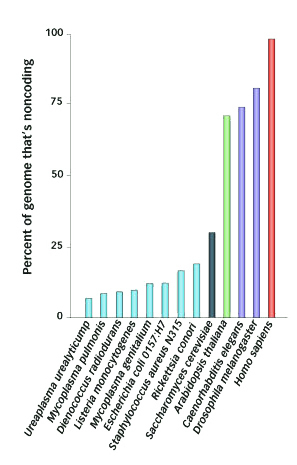
The amount of noncoding DNA is plotted for various species with sequenced genomes. The bars in light blue are bacterial species, the dark gray is brewer’s yeast, the green bar is a plant, the purple bars are two invertebrates, and the orange bar is humans. Image: Science News
A decade ago, human genetics seemed straightforward, writes Elizabeth Pennisi in the Insights of the Decade article "Shining a Light on the Genome’s 'Dark Matter'":
"DNA told the body how to build proteins. The instructions came in chapters called genes. Strands of DNA’s chemical cousin RNA served as molecular messengers, carrying orders to the cells' protein factories and translating them into action. Between the genes lay long stretches of "junk DNA," incoherent, useless, and inert."
It turned out to be nowhere near that simple. Gene regulation is now known to be a complex process governed by regulatory DNA that lies between the genes. RNAs of all shapes and sizes are not just messengers but powerful players in how genomes operate. And chemical alterations called epigenetic factors can influence the genome across generations without changing the DNA sequence itself.
The simple schema started unraveling as soon as the human genome was published in 2001, with protein-coding regions making up only 1.5% of the genome. It seemed unlikely that the rest of our DNA was just junk.
JGI's research in comparative genomics, starting with pufferfish, soon revealed that much of the noncoding DNA had been conserved over eons of evolution, and therefore must perform a useful function. JGI's comparisons of the human and mouse genomes and testing of transgenic mouse embryos then revealed the regulatory function of the noncoding DNA. The leaders of these studies included Eddy Rubin, director of both JGI and Berkeley Lab’s Genomics Division; JGI Genomic Technologies Department Head Len Pennacchio; and JGI Plant Program Head Daniel Rokhsar. Data analyses were performed at JGI and NERSC. Thanks to the efforts of these researchers and many others around the world, the term "junk DNA," though still popular, is now recognized as a misnomer and is most often used ironically.
Tiny Time Machines

Neanderthals are the closest hominid relatives of modern humans. The two species co-existed in Europe and western Asia as recently as 30,000 years ago. DNA analysis has revealed that some Neanderthals had red hair and pale skin. Image: Joe McNally, National Geographic Society.
The Insights of the Decade article "Tiny Time Machines Revisit Ancient Life" by Ann Gibbons relates how the study of ancient life no longer depends solely on structural analysis of fossils, bones, teeth, and shells. "'Biomolecules' such as ancient DNA and collagen can survive for tens of thousands of years and give important information about long-dead plants, animals, and humans," Gibbons writes.
Over the past decade, these molecules have shown that some Neanderthals had red hair and pale skin, and that a Sinosauropteryx dinosaur had a chestnut-colored downy tail. The amino acid sequence of collagen from a dinosaur more closely resembles that of living birds than that of reptiles, confirming the modern theory of bird evolution. DNA has even shown that a few Neanderthals interbred with our ancestors. And recently a previously unknown species of human was identified by its DNA alone.
JGI researchers led by Rubin were among the pioneers of paleogenomics, as this field of research is coming to be known. Until recently, many experts doubted that useful genetic information could be recovered from ancient fossils; in the 1980s such efforts had failed because of contamination with bacterial and modern human DNA. But taking advantage of improved technologies, in 2005 JGI researchers and collaborators from other institutions published DNA sequences from Pleistocene cave bear bones more than 40,000 years old. And the next year they were among the first to sequence nuclear DNA from a Neanderthal fossil.
NERSC supported these studies by providing data storage and management services to JGI. In 2005 JGI's Production Genome Facility was on the verge of generating data faster than they could find somewhere to store the files. NERSC’s High Performance Storage System (HPSS) provided ample data storage capacity; and NERSC collaborated with JGI to improve the data handling capabilities of the genome sequencing and data distribution processes, thus improving the reliability of data storage, making data retrieval easier, and boosting the efficiency of the entire process.
Neighborly Microbes
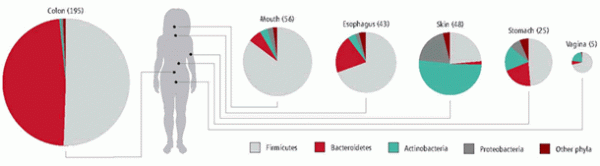
The roster of bacteria varies among body sites. Image: Science Magazine, adapted from L. Dethlefsen et al., Nature 449 (18 OCTOBER 2007).
"This past decade has seen a shift in how we see the microbes and viruses in and on our bodies," writes Elizabeth Pennisi in the Insights of the Decade article "Body's Hardworking Microbes Get Some Overdue Respect." She continues:
"There is increasing acceptance that they are us, and for good reason. Nine in 10 of the cells in the body are microbial. In the gut alone, as many as 1000 species bring to the body 100 times as many genes as our own DNA carries. A few microbes make us sick, but most are commensal and just call the human body home. Collectively, they are known as the human microbiome."
Recent studies have shown that those gut microbes help us harvest more energy and nutrients from our food and even promote digestive health. Other studies have found that our immune system needs friendly bacteria and viruses to develop and function properly. Exploration of the human microbiome is just beginning, but researchers hope to discover ways to manipulate our bodily ecosystem to improve health and combat illnesses.
The Human Microbiome Project, funded by the National Institutes of Health, aims to promote these kinds of discoveries by characterizing the microbial communities found at several different sites on the human body, including nasal passages, oral cavities, skin, gastrointestinal tract, and urogenital tract, and to analyze the role of these microbes in human health and disease.
JGI Chief Informatics Officer and Associate Director Victor Markowitz and Metagenome Program Head Nikos Kyrpides lead the effort to maintain the data catalog associated with the Human Microbiome Project. Markowitz is also head of CRD’s Biological Data Management and Technology Center, which collaborated with JGI to develop one of the analysis tools that makes microbiome research possible: the Integrated Microbial Genomics with Microbiome samples (IMG/M) system, which enables comparative analysis for the study of metagenomes—the collective genetic material of a given microbiome. First released in 2006, IMG/M contains millions of annotated microbial gene sequences, recovered from wild varieties of microbial communities.
A Recipe for the Cosmos

This image of the microwave sky was synthesized using data spanning the range of light frequencies detected by the Planck CMB mission. Image: ESA, HFI, and LFI consortia.
Genomics is not the only field that was transformed in the last decade. Cosmology, writes Adrian Cho in "A Recipe for the Cosmos," has advanced “from a largely qualitative endeavor to a precision science with a standard theory that provides little wiggle room for other ideas,” since it has "only a half-dozen adjustable parameters."
The latest recipe for the cosmos consists of 4.56% ordinary matter, 22.7% dark matter, and 72.8% dark energy. Even though scientists don’t know what two of the three ingredients actually consist of, they do know the precise proportions. What makes this level of detail possible is extraordinarily precise measurements of the cosmic microwave background (CMB), the afterglow of the Big Bang. Cho explains:
The temperature of the CMB varies by about one part in 100,000 from point to point on the sky, with the hotter spots corresponding to the denser regions in the primordial universe. By measuring the distribution of the spots’ sizes and fitting it with their theoretical model, scientists can probe the interplay and amounts of ordinary and dark matter in the early universe. They can also measure the geometry of space. That allows them to deduce the total density of energy and matter in the universe and infer the amount of dark energy.
Since its first allocation of supercomputer resources at Berkeley Lab in 1997, NERSC has become the world leader in providing high performance computing support for CMB data analysis. The balloon-borne missions BOOMERANG and MAXIMA were the first CMB experiments to ever use supercomputing resources for data analysis. (NERSC also supported the analysis of the Supernova Cosmology Project’s Type 1a supernova data, which led to the discovery of dark energy in 1998.)
As the CMB community started incorporating supercomputers into their scientific process, they also had to develop new data analysis methods to manipulate an entire dataset simultaneously and coherently. That’s when Berkeley Lab’s Julian Borrill, Andrew Jaffe, and Radek Stompor stepped in and developed MADCAP, the Microwave Anisotropy Dataset Computational Analysis Package. Researchers used this software to create the maps and angular power spectra of the BOOMERANG and MAXIMA data that confirmed that the universe is geometrically flat, as predicted by the inflation theory of cosmic evolution.
Borrill now co-leads the Computational Cosmology Center, a collaboration between CRD and the Lab's Physics Division to pioneer algorithms and methods for optimizing CMB research on cutting-edge supercomputer technology.
"When NERSC first started allocating supercomputing resources to the CMB research community in 1997, it supported half a dozen users and two experiments. Almost all CMB experiments launched since then have used the center for data analysis in some capacity, and today NERSC supports around 100 researchers from a dozen experiments," says Borrill.
Currently the biggest CMB project at NERSC is the Planck mission, which on January 11, 2011 released a new catalog of data from its initial maps of the entire sky. Planck is observing the sky at nine wavelengths of light, ranging from infrared to radio waves. The result is a windfall of data on known and never-before-seen cosmic objects. The mission will provide the cleanest, deepest, and sharpest images of the CMB ever made, and NERSC has optimized its computing and storage systems to handle that data efficiently.
For further reading:
- Whole-Genome Shotgun Assembly and Analysis of the Fugu rubripes Genome
- Ancient Cave Bear DNA Successfully Sequenced
- Neanderthal Genome Sequencing Yields Surprising Results and Opens a New Door to Future Studies
- Data Acquisition and Coordination Key to Human Microbiome Project
- BOOMERANG Antarctic Balloon Flight Sees a Flat Universe: NERSC Supercomputer Analysis Crucial to the Result
- Mapping the Universe to the Beginning of Time
- NERSC Continues Tradition of Cosmic Microwave Background Data Analysis with the Planck Cluster
- Planck Mission Peels Back Layers of Universe
About Computing Sciences at Berkeley Lab
High performance computing plays a critical role in scientific discovery. Researchers increasingly rely on advances in computer science, mathematics, computational science, data science, and large-scale computing and networking to increase our understanding of ourselves, our planet, and our universe. Berkeley Lab’s Computing Sciences Area researches, develops, and deploys new foundations, tools, and technologies to meet these needs and to advance research across a broad range of scientific disciplines.







 Instagram
Instagram YouTube
YouTube