Decoding the Molecular Mysteries of Photosynthesis
February 14, 2014
Contact: Kathy Kincade, +1 510 495 2124, kkincade@lbl.gov
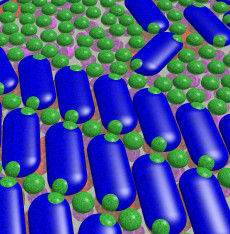
Two protein assemblies in a plant cell's chloroplasts -- Photosystem II (blue and red) and light-harvesting complex II (green and purple) -- are key to initiating photosynthesis. This visualization illustrates how the proteins organize themselves to make this happen. (Image by Anna Schneider, UC Berkeley, and Lester Hodges, Berkeley Lab)
At first glance, photosynthesis seems elegant in its simplicity. Only three elements – light, water and carbon dioxide (CO2) – are needed for plants and other organisms to convert light into chemical energy, all in a matter of minutes.
But upon closer inspection, the process is much more complicated. How, for example, does the plant know how much sunlight, water or CO2 it needs at any given point in time? And what if the system breaks – how does the plant repair it?
Scientists are working to answer these questions to enable the development of artificial photosynthesis systems that can generate “green” alternatives to fossil fuels. But first they must pinpoint what happens at the molecular level that allows photosynthesis to occur in the first place.
Toward that end, simulations by researchers from the University of California, Berkeley (UC Berkeley) and the Lawrence Berkeley National Laboratory (Berkeley Lab) using supercomputers at the Department of Energy’s National Energy Research Scientific Computing Center (NERSC) have shed new light on the inner workings of the photosynthetic process. The team’s paper appeared as the cover story in the September 2013 Biophysical Journal.
“In artificial photosynthesis the systems can’t repair themselves if they break and they can’t tune their efficiency up and down based on what the system needs,” explained lead author Anna Schneider, a recent graduate of the Biophysics Graduate Group at UC Berkeley. “But these are things that plants can do, and we are trying to figure out the physics that underlie that so that someone down the line can build something that takes advantage of this.”
Finding Order in Nature’s ‘Tiny Green Poker Chips’
All photosynthesis occurs in a plant’s chloroplasts--organelles contained within the plant cells. A typical plant cell can contain up to 50 chloroplasts. A chief component of the chloroplasts is the thylakoid membrane, which contains discs of protein assemblies arranged in stacks (grana) that resemble towers of tiny green poker chips. In recent years much photosynthesis research has focused on teasing apart one of these protein assemblies: Photosystem II (PSII).
“PSII is a large complex containing dozens of proteins and hundreds of chlorophylls and other pigments,” she said. “PSII catalyzes the first step in the photosynthetic electron transport chain, setting an upper limit for overall photosynthetic efficiency and productivity.” Each chloroplast contains thousands of PSII complexes, she noted.
Photosynthetic efficiency relies on precise spatial organization of PSII and its associated light-harvesting complex II (LHCII), both of which are concentrated in the grana stacks. But how the proteins know to arrange themselves is unclear, according to Schneider and her co-author, Phillip Geissler, professor of chemistry at UC Berkeley. This makes it difficult to determine exactly how they regulate a plant’s light harvesting and energy conversion efficiencies.
“The grana protein organization affects the efficiency of photosynthesis overall,” Schneider said. “Our goal was to develop a physical understanding of why the proteins are organized the way they are.”
The Thermodynamics of Ice and Water
To do this they created a nanoscale computational model of LHCII and PSII and used it to analyze the protein organization. The model is simple enough to access large length scales, yet rich enough to capture structural motifs such as inter-membrane correlations, Schneider explained. In addition, it can predict thermodynamic phase behavior that could affect photosynthetic function.
“If you think about water, you can have liquid, ice and vapor, and those are each different thermodynamic phases,” Schneider explained. “But if you have cubes floating in water, you have two phases coexisting at the same time and interfacing with one another. But when you get down to the nanoscale, it is hard to tell if what you have is an ice cube or a few water molecules that happen to be next to each other in a way that looks like ice. “
The computer model was designed to enable researchers to make this distinction, and to do so with a very small set of variables.
“That was the point of our simulations: to pin down that it’s not just that the PSII and LHCII happen to be next to each other in a way that looks like a crystal—it is a true thermodynamic crystal with true separate phases,” Schneider said.
This discovery means that scientists can apply everything they already know about the thermodynamics of ice and water to what’s going on inside plants, she added.
For example, using umbrella sampling and running Monte Carlo simulations on NERSC’s Hopper supercomputer, she and her colleagues showed that the simulated PSII-LHCII arrays are evidence of a true crystalline phase that coexists with a fluid phase within the chloroplast model. As a result, small changes in protein density or interactions could lead to dramatic shifts in photosynthetic behavior, Schneider noted.
“We found that there is a barrier for changing between those two phases and that by changing what is going on inside the plants just a tiny bit, you can get very dynamic results in terms of the regulatory process,” she explained. “This is something the plants can take advantage of when changing their efficiencies.”
Customized Visualization Tools
Being able to visualize their analysis along the way was critical, Schneider emphasized.
“The multilayered structure of our configurations was difficult to visualize clearly using traditional software,” she explained. Instead, she used a tool that she developed while taking a data visualization class from Maneesh Agrawala, professor of Electrical Engineering and Computer Science at UC Berkeley. She then collaborated with Lester Hedges, a post-doc working in Berkeley Lab’s Molecular Foundry, to create 3D images of the data using POV-Ray, a free software visualization tool. One of these images was featured on the cover of Biophysical Journal.
“It may not look like there is that much data in some of the figures (in the paper), but it actually took several hundred cores on Hopper running for several weeks to develop that data,” Schneider said.
She is now working to expand the model to address other unknowns about photosynthesis at the molecular level.
“This paper was focused on looking inside the grana at PSII and LHCII, but there are other questions that are important for understanding things like how the system repairs itself when it breaks,” she said. “There are other components we need to understand in order to analyze the physics.”
About Computing Sciences at Berkeley Lab
High performance computing plays a critical role in scientific discovery. Researchers increasingly rely on advances in computer science, mathematics, computational science, data science, and large-scale computing and networking to increase our understanding of ourselves, our planet, and our universe. Berkeley Lab’s Computing Sciences Area researches, develops, and deploys new foundations, tools, and technologies to meet these needs and to advance research across a broad range of scientific disciplines.







 Instagram
Instagram YouTube
YouTube