Data Processing
In order to move from raw data to scientific knowledge, data must go through a series of transformations that require a combination of software tools, compute infrastructure, and storage systems to generate the necessary data pipelines. We work closely with science teams to develop these workflows, connecting sensor networks, edge computing devices, and high-end DOE experimental facilities to large-scale compute and storage infrastructure.
With one of the largest supercomputers at the DOE complex, Berkeley Lab is a prime location for data processing in science. Surrounding this resource, our researchers have developed vast arrays of data analysis and data processing tools and algorithms. These tools can extract features from noisy data, calibrate large scientific instruments, generate tomographic images from different forms of projection data, assemble millions of DNA fragments into coherent structures, identify anomalies in data streams, recommend actions for on-going experiments, and more.
Projects
Jupyter
Jupyter is an essential component of the modern interactive data ecosystem. Berkeley lab staff are working to enable tools and services that can bridge the gap between interactive notebook computing, and scalable high performance computing systems. Our work allows users to perform exploratory data analytics and visualization of data stored on the NERSC Global File System or in databases at NERSC; guide machine learning through distributed training, hyperparameter optimization, model validation, prediction, and inference; and manage workflows involving complex simulations and data analytics through the batch queue. Contacts: Rollin Thomas, Shreyas Cholia
ExaBiome
The ExaBiome project is developing HipMer and MetaHipMer, the first high-quality, end-to-end de novo genome and metagenome assemblers designed for extreme scale analysis. Both assemblers are PGAS applications, and the main software dependencies are the UPC language and UPC++ library, both of which use Berkeley Lab’s GASNet-EX library for communication. The high performance of these assemblers is based on several novel algorithmic advancements attained by leveraging the efficiency and programmability of the one-sided communication capabilities of the Unified Parallel C (UPC), and RPC calls from the UPC++ Library, including optimized high-frequency k-mer analysis, communication avoiding de Bruijn graph traversal, advanced I/O optimization, and extensive parallelization across the numerous and complex application phases. Contact: Katherine Yelick (Yelick on the Web)
Superfacility
The Superfacility model is designed to leverage HPC for experimental science. It is more than simply a model of connected experiment, network, and HPC facilities; it encompasses the full ecosystem of infrastructure, software, tools, and expertise needed to make connected facilities easy to use. The Berkeley Lab Superfacility Project was initiated in 2019 to coordinate work being performed at the Lab in support of this model and to provide a coherent and comprehensive set of science requirements to drive existing and new work. There is a pressing need for a coherent computing infrastructure across national facilities, and our Superfacility Project is a unique model for success in tackling the challenges that will be faced in hardware, software, policies, and services across multiple science domains. The lessons we learned during this project provide a valuable example for future large, complex, cross-disciplinary collaborations. Contact: Deborah Bard
AmeriFlux
AmeriFlux, launched in 1996, is a network of PI-managed sites measuring ecosystem CO2, water, and energy fluxes in North, Central and South America. In 2012 DOE established the AmeriFlux Management Project (AMP) at Berkeley Lab to support the broad AmeriFlux community and the AmeriFlux sites (currently more than 110 active sites). AMP collaborates with AmeriFlux scientists to ensure the quality and availability of the continuous, long-term ecosystem measurements necessary to understand these ecosystems and to build effective models and multisite syntheses. Data. Deb Agarwal, who leads Berkeley Lab Computing Sciences’ Scientific Data Division, is the AMP Data Team Lead. Contact: Deborah Agarwal
Saturn
Plain-text Python notebooks with checkpointing. Contact: Dmitriy Morozov
TECA
ECA is a collection of climate analysis algorithms geared toward extreme event detection and tracking implemented in a scalable parallel framework. The code has been successfully deployed and run at massive scales on current DOE supercomputers. TECA's core is written in modern C++ and exploits MPI + X parallelism where X is one of threads, OpenMP, or GPUs. The framework supports a number of parallel design patterns including distributed data parallelism and map-reduce. While modern C++ delivers the highest performance, Python bindings make the code approachable and easy to use. Contact: Burlen Loring
Prototyping a monitoring system of global wetland CH4 emissions with machine learning and satellite remote sensing
This project will provide a radical solution to the big challenges of monitoring wetland CH4 using a novel machine learning approach. We will deliver the first high-resolution, multi-decadal gridded product of global wetland CH4 flux. It will improve our understanding of wetland CH4 emission, which is the most important natural CH4 source and a great contributor to global warming. Contact: Qing Zhu
pyHPC for microCT
This project aims at accelerating microstructural analytics with Dask for volumetric X-ray images using HPC systems, such as NERSC. Developments under this initiative enable many light sources users to have improved and high level access to image processing capabilities that run in parallel, which are some of the key elements to a SuperFacility. Contact: Dani Ushizima
Spin
Spin is a container-based platform at NERSC designed to deploy your science gateways, workflow managers, databases, API endpoints, and other network services to support scientific projects. Services in Spin are built with Docker containers and can easily access NERSC systems and storage. NERSC users are able to share the data and tools that enable science through the center’s science gateways. Contacts: Cory Snavely, Annette Greiner, Daniel Fulton
News
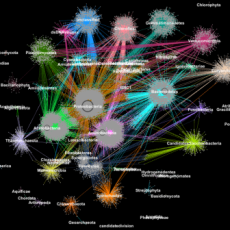
A Game Changer: Metagenomic Clustering Powered by HPC
A team of researchers from Berkeley Lab's CRD and JGI took one of the most popular clustering approaches in modern biology—the Markov Clustering algorithm—and modified it to run quickly, efficiently and at scale on distributed-memory supercomputers. Read More »
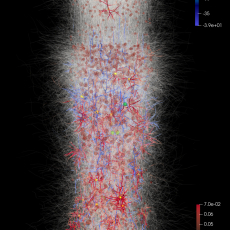
Neuroscience Simulations at NERSC Shed Light on Origins of Human Brain Recordings
Using simulations run at NERSC, a team of researchers at Berkeley Lab has found the origin of cortical surface electrical signals in the brain and discovered why the signals originate where they do. Read More »

Autonomous Discovery: What’s Next in Data Collection for Experimental Research
A Q&A with CAMERA's Marcus Noack on this emerging data acquisition approach and a related virtual workshop for users from multiple research areas. Read More »
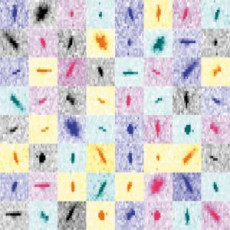
Novel Data Processing Technique Aids XFEL Protein Structure Analysis
Scientists use X-ray crystallography to determine the three-dimensional structures of proteins. A new data processing approach is enhancing the outcomes. Read More »

Physics Data Processing at NERSC Dramatically Cuts Reconstruction Time
In a recent demonstration project, physicists from Brookhaven National Laboratory and Berkeley Lab used the Cori supercomputer to reconstruct data collected from a nuclear physics experiment, an advance that could dramatically reduce the time it takes to make detailed data available for scientific discoveries. Read More »

Supercomputing Pipeline Aids DESI’s Quest to Create 3D Map of the Universe
The Dark Energy Spectroscopic Instrument is combining high-speed automation, high-performance computing, and high-speed networking to produce the largest 3D map of the universe ever created. Read More »







 Instagram
Instagram YouTube
YouTube